IF Spatial Dots
Perform spatial analysis of dot-like signals (e.g., FISH, RNAscope) within nuclei, quantifying nuclei area, dots per nucleus, and dot distribution by direction and proximity.
FISH, cell culture, RNAScope, spatial analysis, proximities, nuclei orientation

The IF Spatial Dots App allows in-depth spatial analysis of dot-like stainings (FISH, RNAScope etc.) within nuclei. It outputs the number and area (µm2) of nuclei, number of detected dots per nuclei, number/% of dots per nuclei direction and proximity.
Irina Milisav
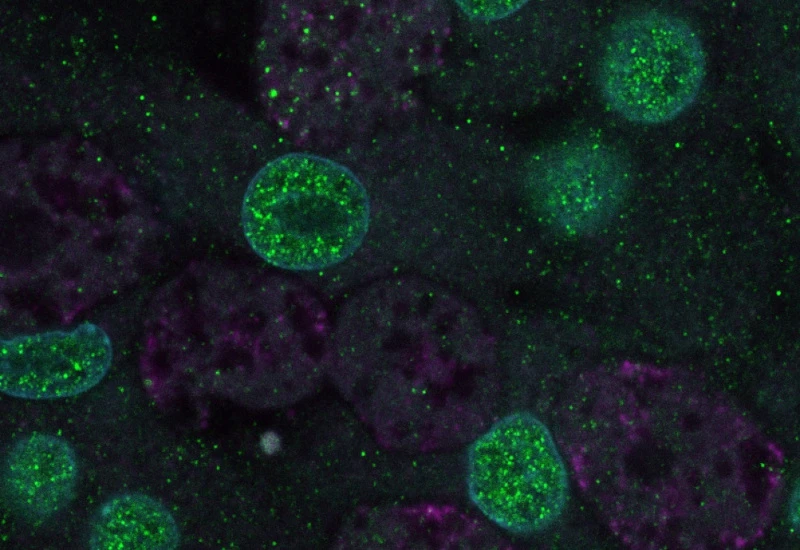
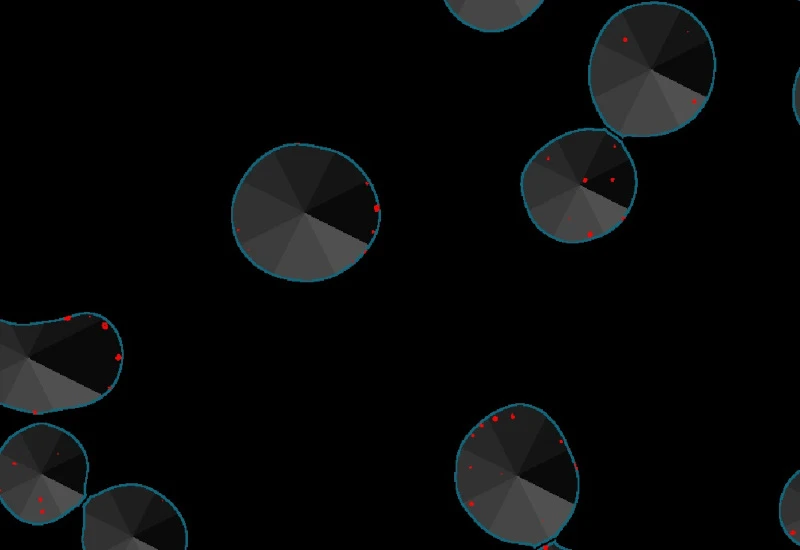
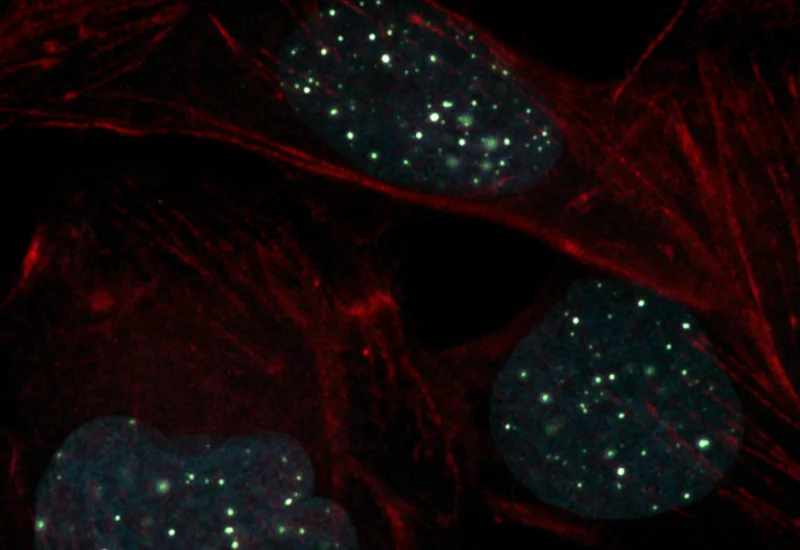
Original image
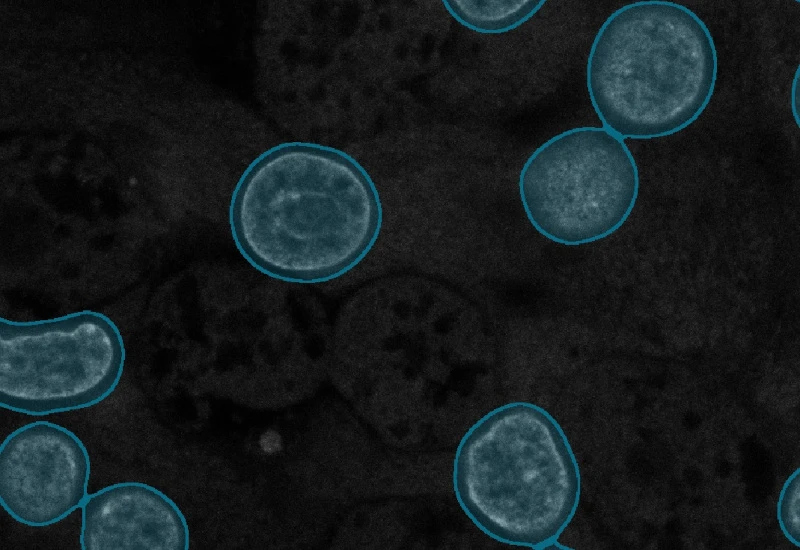
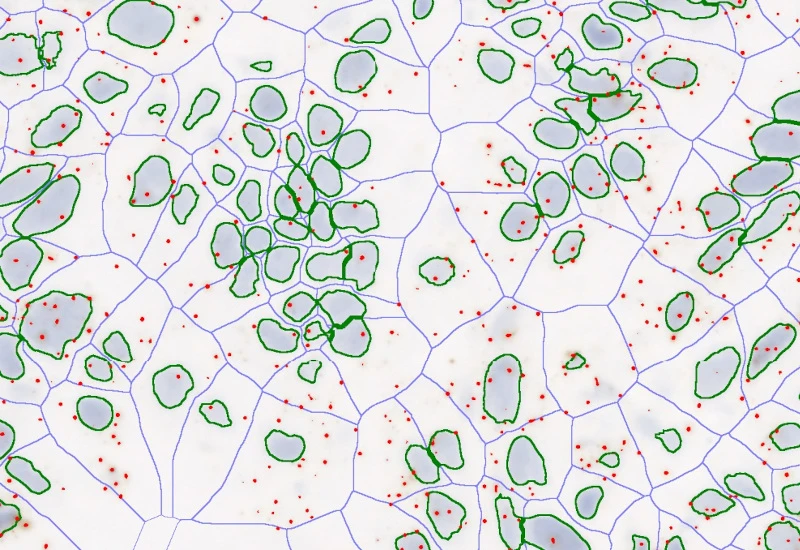
Nuclei detection
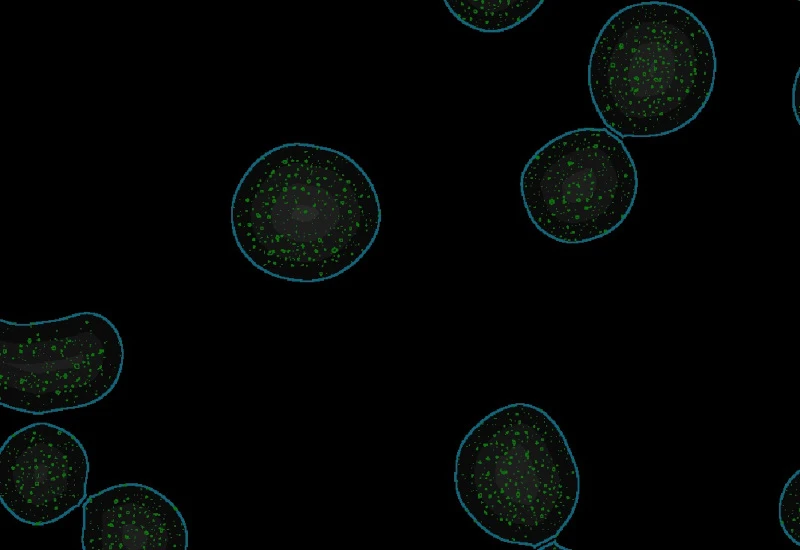
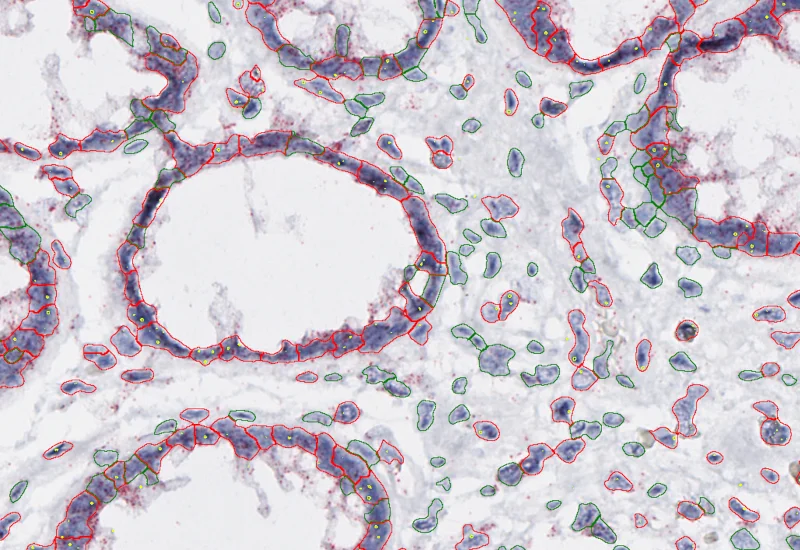
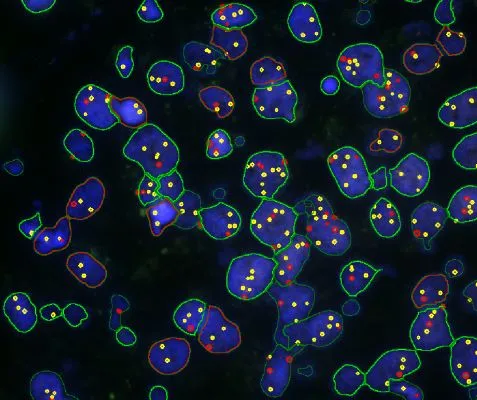
Proximity map of detected dots

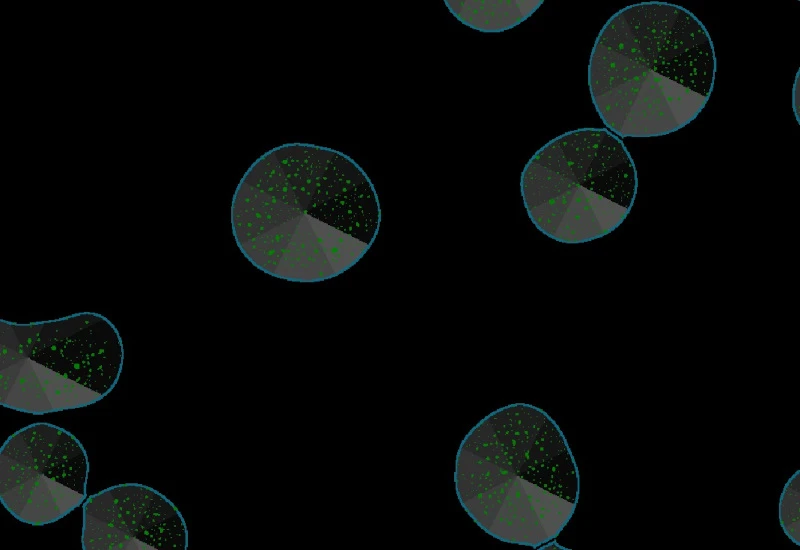
Direction estimation of detected dots
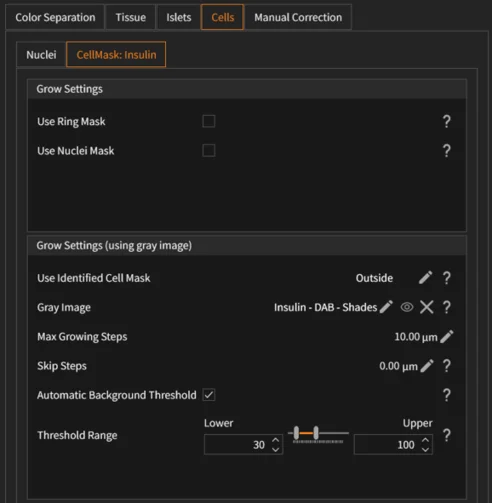
The IF Spatial Dots App allows users to adjust nuclei detection, proximity mapping, and dot detection parameters per fluorescence channel:
- Nuclei detection: Select input image, adjust nuclei size, define background threshold value, and remove nuclei by area or intensity.
- Proximity areas: Define custom distance values (µm or pixels) to generate concentric proximity zones inside nuclei; add, delete, or modify distance lists.
- Spatial directions: Analyze dot distribution across 8 directional sectors within each nucleus.
- Dot detection (per channel): Select input image and events mask, configure dot size range, window size range (kernel size), sensitivity, and background value to control detection stringency.
- Manual correction: Add, remove, split, or merge nuclei using interactive drawing tools.
- Overlay validation: Review nuclei masks, proximity maps, directional sectors, and detected dots before exporting results.
The app provides quantitative measurements at nuclei and spatial dot levels:
- Nuclei metrics: Area (µm²) per nucleus and total number of nuclei.
- Dots per nucleus: Number of detected dots per nucleus and total dot count per channel.
- Directional analysis: Number and area (µm²) of dots in each of 8 spatial directions (Dir 1–8) per nucleus and total per sample.
- Proximity analysis: Number and area (µm²) of dots within defined distance ranges (e.g., 0–5 µm, 5–10 µm, etc.) inside nuclei.
- Center localization: Number of dots located inside or outside the nucleus center.
- Raw data access: Per-nucleus measurements available in the Raw Data table.
- Statistics export: ROI/sample-based results generated via the Statistics Report tool.
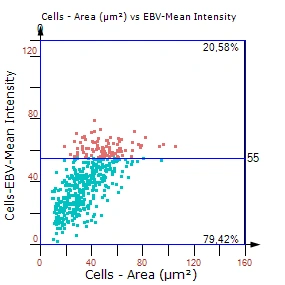
Fig.1: Example plot
Customer Publication
12 Oct, 2021
Tissue Cytometry in HIV Research
Application Note
14 Feb, 2023
Case Study: Analysis of FISH Using Tissue Cytometry
Blog Post
15 Feb, 2023
Applications of AI in Cell Segmentation
We support the following file formats:
- TissueFAXS (aqproj)
- StrataFAXS II (vmic)
- PreciPoint (vmic, gtif)
- Generic BigTIFF Import
- Support for multipage BigTIFF files
- OME-TIFF
- JPEG, PNG, BMP, TIFF
- Zeiss (czi)
- Hamamatsu NanoZoomer (ndpi)
- Aperio (svs)
- Leica (scn)
- 3D HISTECH Pannoramic
- Mirax (mrxs)
- Olympus (vsi)
- More slide scanners to be added!
Related Apps
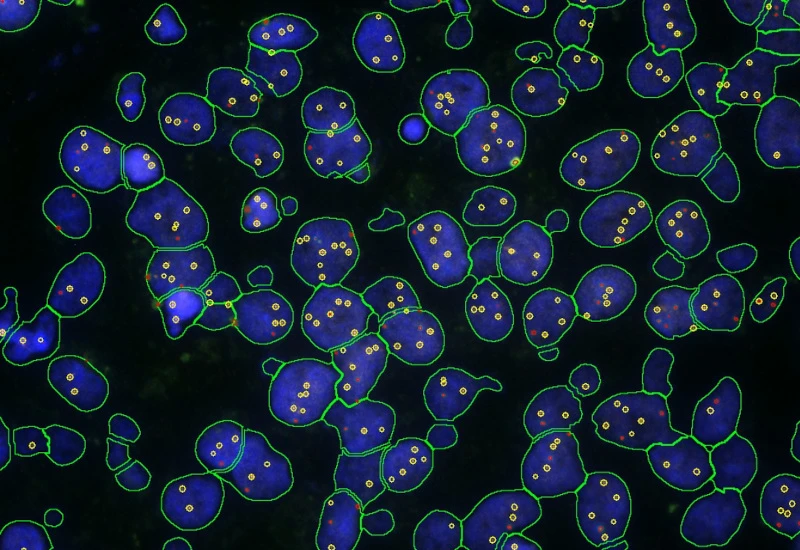
IF Dots
Detect dots-stainings per cell in up to four markers, segment cellular compartments, measure up to 20 intensity, statistic, and morphometric parameters, and quantify dot count, mean intensity, total area, intensity sum, and per-dot area/intensity per compartment.
cell culture, breast cancer, fluorescence, HER2
IF Dots Colocalization
Detect dot-like signals (e.g., FISH, RNAscope) in multiple fluorescence channels and quantify dot number, size, and colocalization between markers.
FISH, cell culture, RNAScope, colocalization

Custom App development
Perfectly tailored image analysis solutions for your research.
You have a specific research question that needs to be answered? We offer custom development of image analysis pipelines for specific tasks, be it detection of cellular phenotypes or quantification of tissue structures. After discussing your goals with one of our experts, you will get a ready-to-use App and be a step closer to an impactful publication.