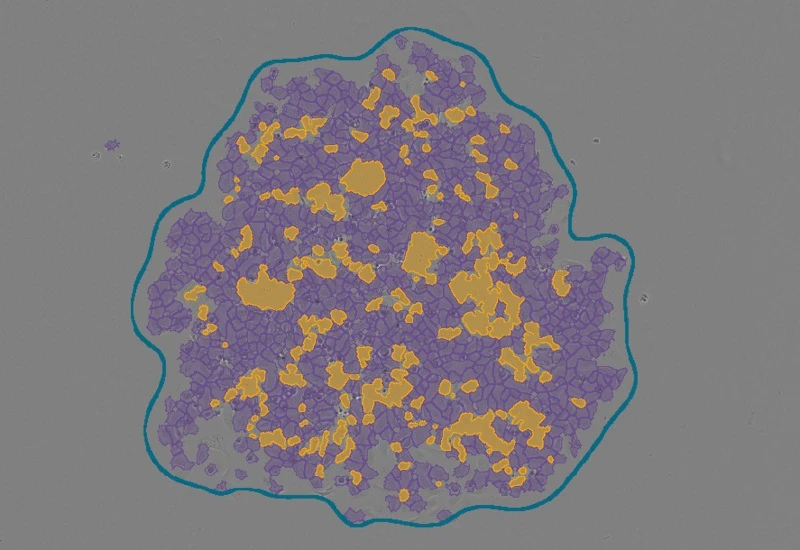
Stem Cell Colony Assay
Identify single cells, cell clusters, and cluster gaps within stem cell colonies, and quantify cluster area, cell area, gap area, and gap-based cluster classification.
stem cells, phase contrast, cell culture, stem cell colony

The Stem Cell Colony App identifes single cells, cell clusters and cluster gabs (no cells) within cell colonies in culture.The output of the App is total cluster area, total cell area, total cluster gabs area and percentage from cluster area. Further it classifiers the clusters based on number of cluster gabs and percentage from each cluster.
Srinivasan S, Kryza T, Bock N, et al. A PSA SNP associates with cellular function and clinical outcome in men with prostate cancer. Nat Commun. 2024;15(1):9587. Published 2024 Nov 6. doi:10.1038/s41467-024-52472-6
Jyotsna Batra, Queensland University of Technology, Brisbane, Queensland (QLD), Australia

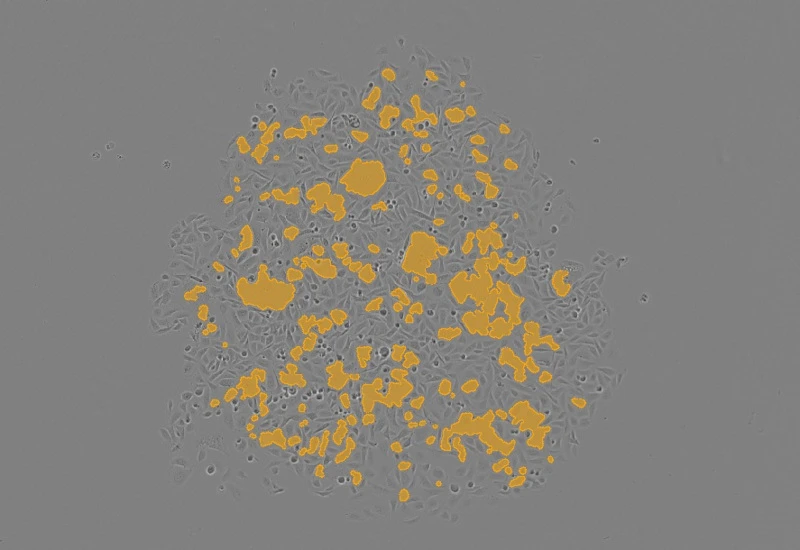
Original Image
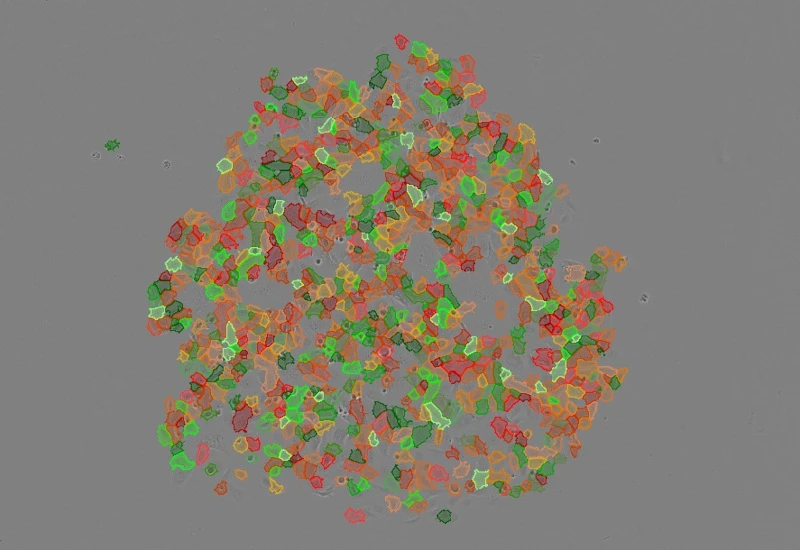

Cell detection

Cluster detection

Cluster gap detection
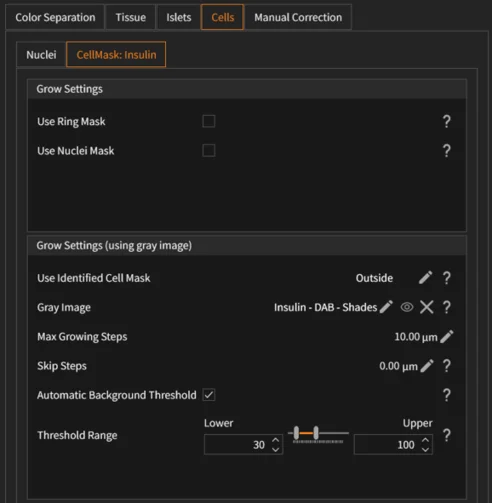
The Stem Cell Colony App allows users to adjust tissue, cell, cluster, and gap detection parameters:
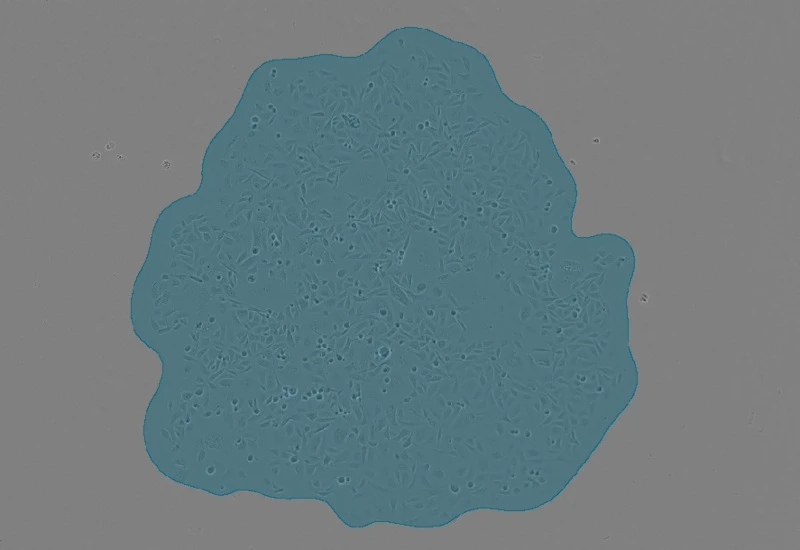
- Tissue detection: Select input image, define threshold value for tissue mask generation, adjust shrink radius, and remove tissue regions based on area (µm²).
- Membrane detection (cells): Configure Master/Membrane input images, remove small or intensity-out-of-range labels, adjust minimum variation sensitivity, kernel width/height, Gaussian filtering (Gauss Std), and area filtering.
- Cell seeds detection: Select input image, adjust threshold for seed extraction, define grow radius to reconstruct full cells, and remove seeds by area (µm²).
- Cluster gaps detection: Adjust grow radius for cluster gaps and remove gaps based on area thresholds.
- Clusters final detection: Select input image, define threshold for cluster segmentation, configure cluster ring exterior radius, and remove clusters by area (µm²).
- Manual correction: Add or delete Tissue, Cell Seeds, or Clusters using interactive drawing tools with adjustable thickness, fill, erase, apply, and cancel options.
- Overlay validation: Review Tissue, Cells, Clusters, and Cluster Gaps masks before exporting results.
The app provides quantitative measurements at colony, cluster, and gap levels:
- Total cluster area: Area (µm²) of detected clusters.
- Total cell area: Area (µm²) of detected cells within colonies.
- Cluster gaps: Total area (µm²) of gaps (cell-free regions) within clusters.
- Gap percentage: Cluster gaps area expressed as percentage of cluster area.
- Cluster classification: Classification of clusters based on number of cluster gaps and percentage of gap area per cluster.
- Statistics export: ROI/sample-based results generated via the Statistics Report tool.
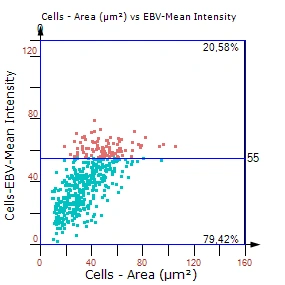
Fig.1: Example plot

Application Note
14 Oct, 2024
Quantitative Analysis of Cultured Cells: High-End Solution for Ready-To-Publish Data
We support the following file formats:
- TissueFAXS (aqproj)
- StrataFAXS II (vmic)
- PreciPoint (vmic, gtif)
- Generic BigTIFF Import
- Support for multipage BigTIFF files
- OME-TIFF
- JPEG, PNG, BMP, TIFF
- Zeiss (czi)
- Hamamatsu NanoZoomer (ndpi)
- Aperio (svs)
- Leica (scn)
- 3D HISTECH Pannoramic
- Mirax (mrxs)
- Olympus (vsi)
- More slide scanners to be added!
Related Apps

Custom App development
Perfectly tailored image analysis solutions for your research.
You have a specific research question that needs to be answered? We offer custom development of image analysis pipelines for specific tasks, be it detection of cellular phenotypes or quantification of tissue structures. After discussing your goals with one of our experts, you will get a ready-to-use App and be a step closer to an impactful publication.


