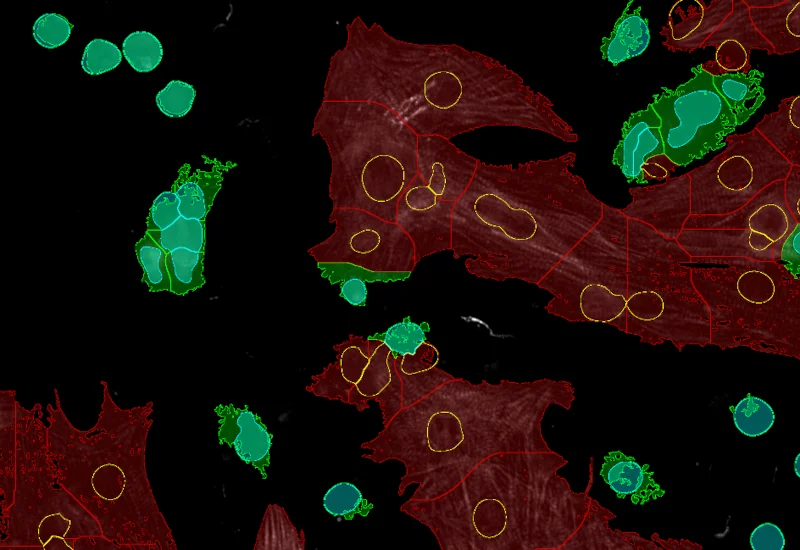
IF Cardio Cell Culture
Detect nuclei and identify cardiomyocytes in IF-stained cell cultures, reconstruct cytoplasmic masks, and quantify cardiomyocyte number, area, and marker intensity.
cardiology, cardiomyocytes, cell culture, fibroblasts, troponin red

The IF Cardio Cell Culture App provides cell segmentation, detection of cardiomyocytes (based an appropriate stain e.g. Troponin Red) and fibroblasts within cultured cardio cells plus one additional marker. The App outputs parameters such as number of cardiomyocytes, fibroblasts and markerpos cardiomyocytes, fibroblasts.
Agatha Ribeiro da Silva, Prof. Jose E. Krieger (Heart Institute, University Soa Paulo)
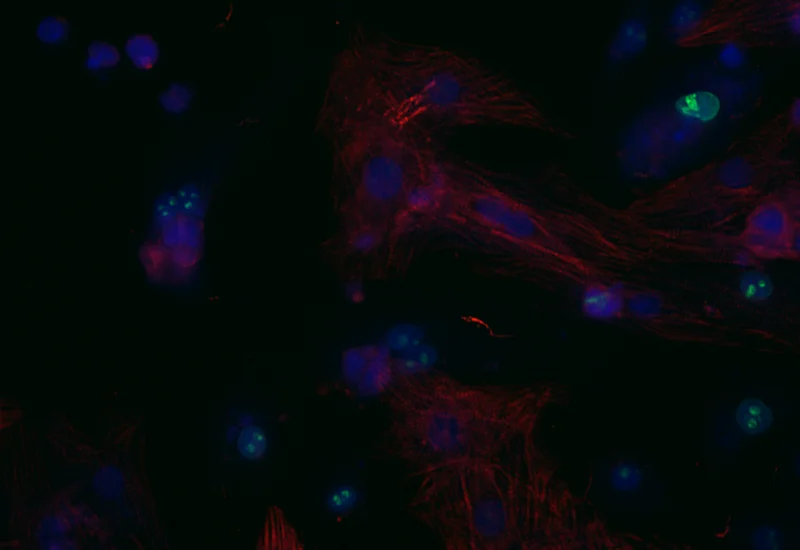
Original Image
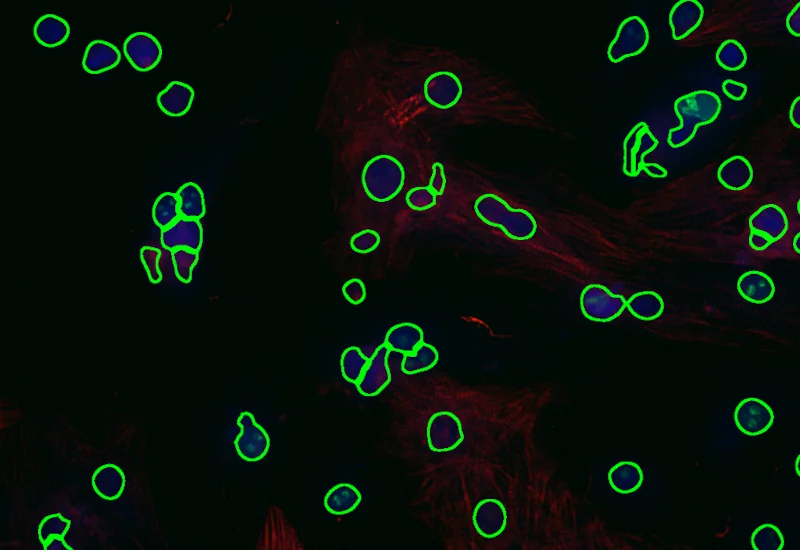
Nuclei detection
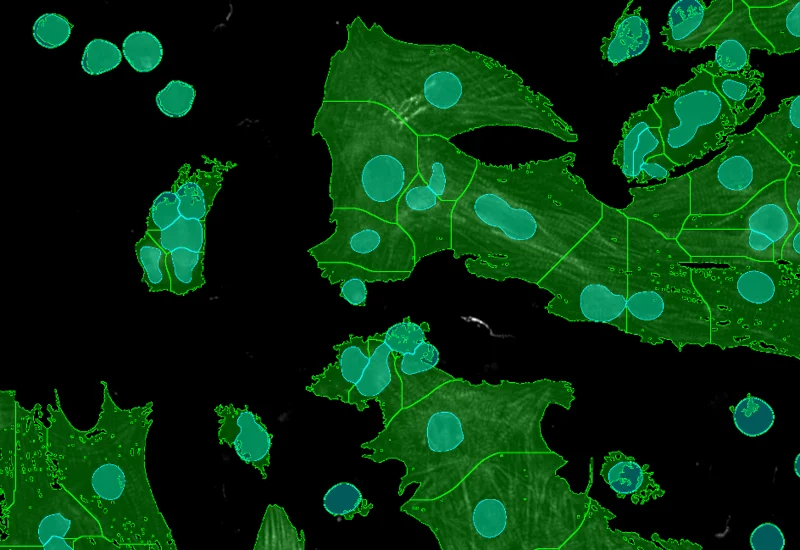
Cellular mask

Cardiomyocytes (red), fibroblasts (green)

Application Note
14 Oct, 2024
Quantitative Analysis of Cultured Cells

Webinar
12 Nov, 2025
TG Academy Webinar - Decoding the language of cancer with SCellBOW

Application Note
14 Oct, 2024
Quantitative Analysis of Cultured Cells
We support the following file formats:
- TissueFAXS (aqproj)
- StrataFAXS II (vmic)
- PreciPoint (vmic, gtif)
- Generic BigTIFF Import
- Support for multipage BigTIFF files
- OME-TIFF
- JPEG, PNG, BMP, TIFF
- Zeiss (czi)
- Hamamatsu NanoZoomer (ndpi)
- Aperio (svs)
- Leica (scn)
- 3D HISTECH Pannoramic
- Mirax (mrxs)
- Olympus (vsi)
- More slide scanners to be added!
Related Apps
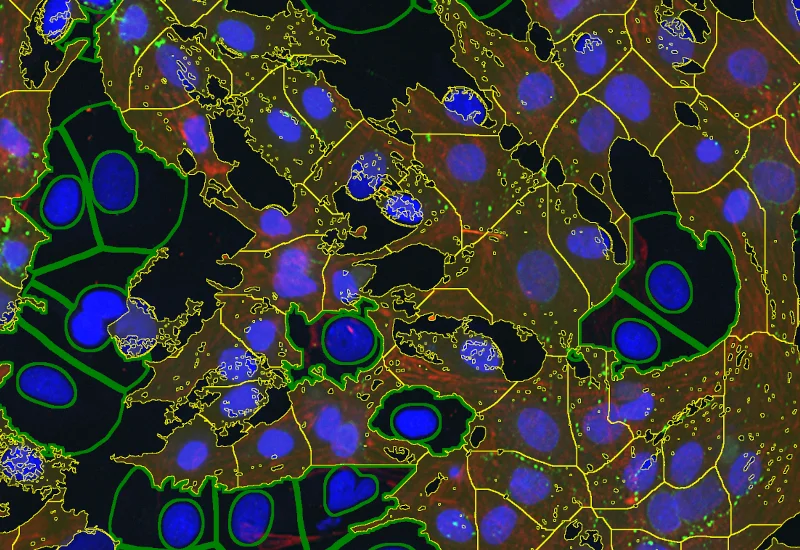
IF Cardio Cell Culture Dots
Segment cultured cardiac cells, detect cardiomyocytes and fibroblasts, and quantify dot markers (CISH/FISH) per cell, including cell counts and dot number, area, and mean intensity.
cardiology, cardiomyocytes, cell culture, fibroblasts, troponin red, FISH
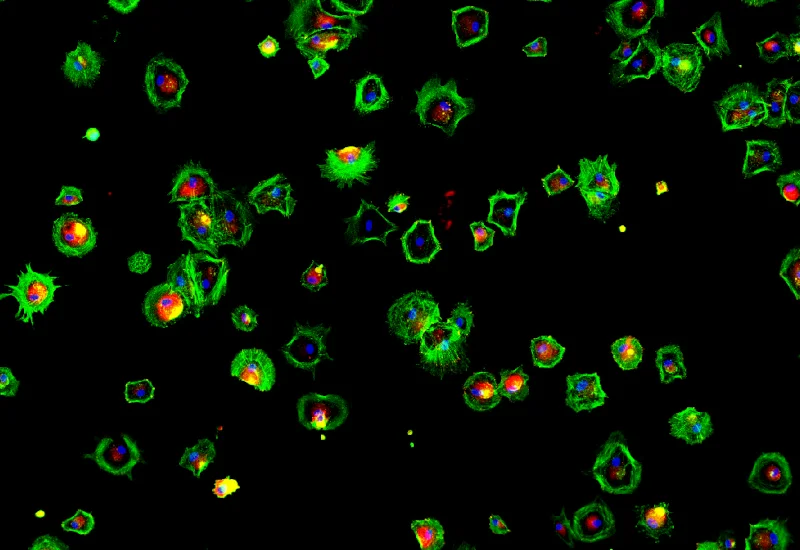
IF Cultured Cells & Substructures
Detect nuclei-based cells, quantify dot markers per cell, and measure cytoskeletal filament density and dot intensity at single-cell level.
FISH, cell culture, cytoskeleton, actin fibres
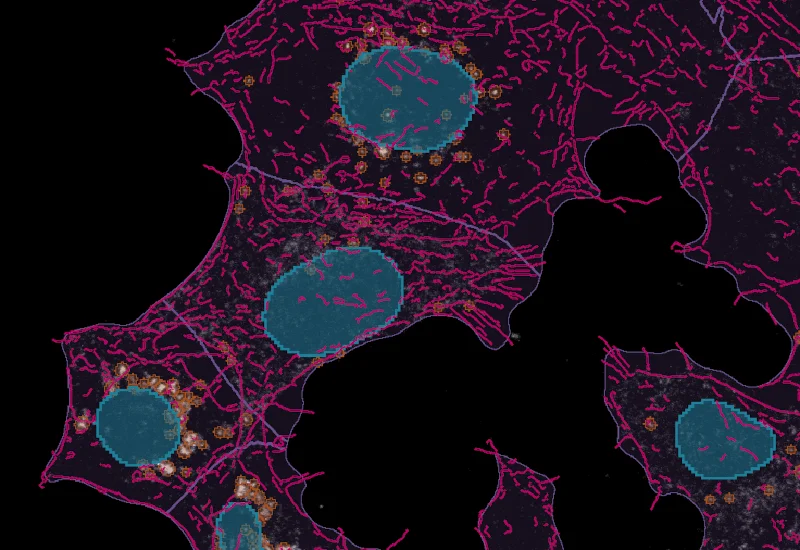
IF Cytoskeleton
Detect cytoskeletal structures by specific stain, identify cytoplasm with additional markers, and export filament count (inside, outside, membrane), filament length, and total filament area.
actin, cytoskeleton, cortical fibres, microfilaments, stress fibres, cell culture, fluorescence

Custom App development
Perfectly tailored image analysis solutions for your research.
You have a specific research question that needs to be answered? We offer custom development of image analysis pipelines for specific tasks, be it detection of cellular phenotypes or quantification of tissue structures. After discussing your goals with one of our experts, you will get a ready-to-use App and be a step closer to an impactful publication.


